Introduction
|
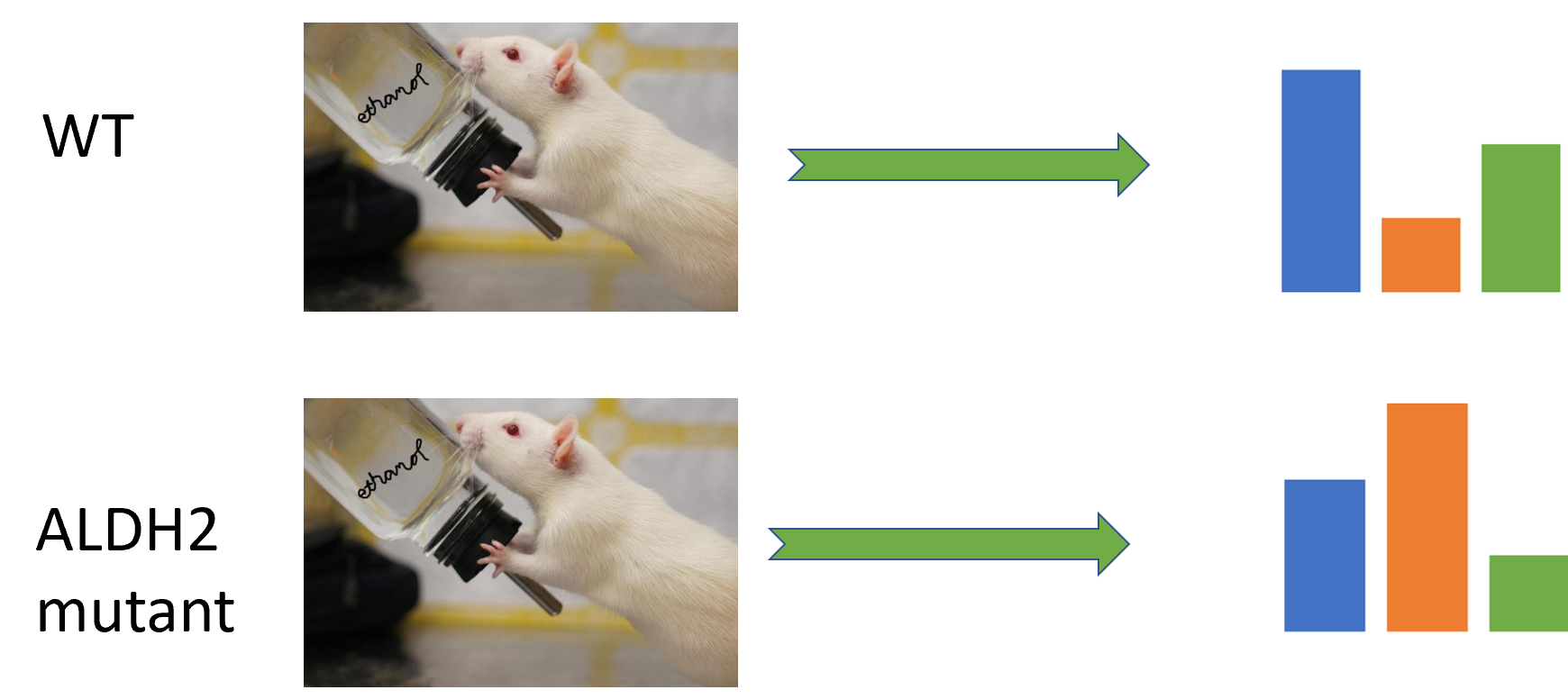
Alcohol intolerance is a widespread autosomal dominant disease, affecting about 8% of the world. People with this disease typically have adverse effects shortly after consuming alcohol. Alcohol flush is known as the immediate reaction for people with this disease. Most people with this disease have a mutation in the ALDH2 protein, particularly known as the E487K mutation. Normally a buildup of the byproducts of alcohol degradation lead to apoptosis or cell death. However, people with this disease have a significantly higher incidence of cancer.
|
ALDH2 gene
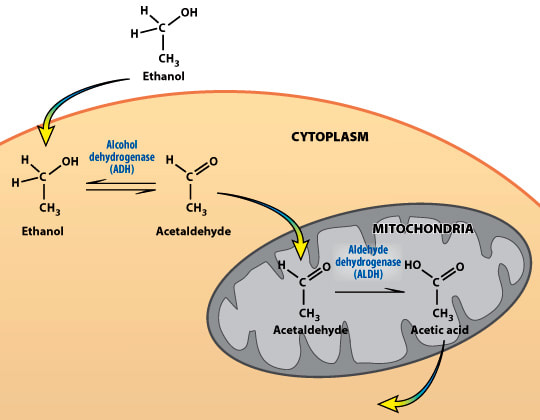
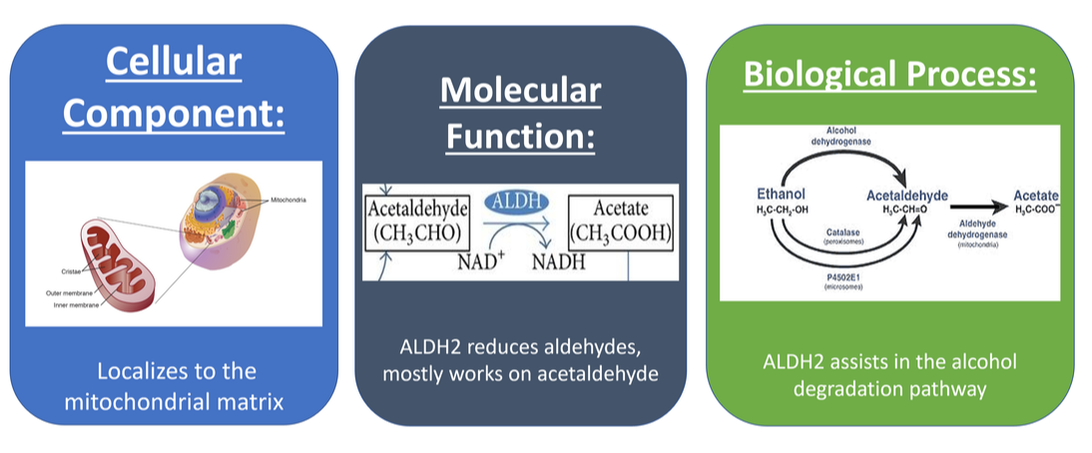
Alcohol intolerance is mostly caused by a mutation in the gene for aldehyde dehydrogenase-2 (ALDH2). This protein's main purpose is to reduce acetaldehyde to acetate. This protein is unlocalized to a specific organelle, and is mostly involved in metabolic functions.
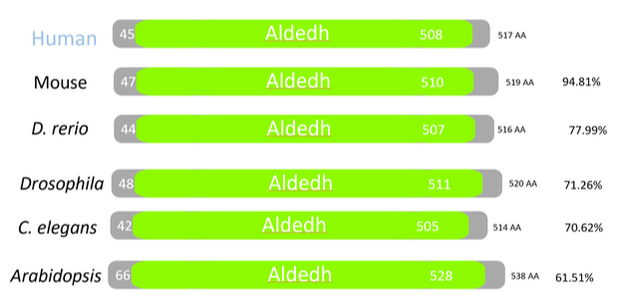
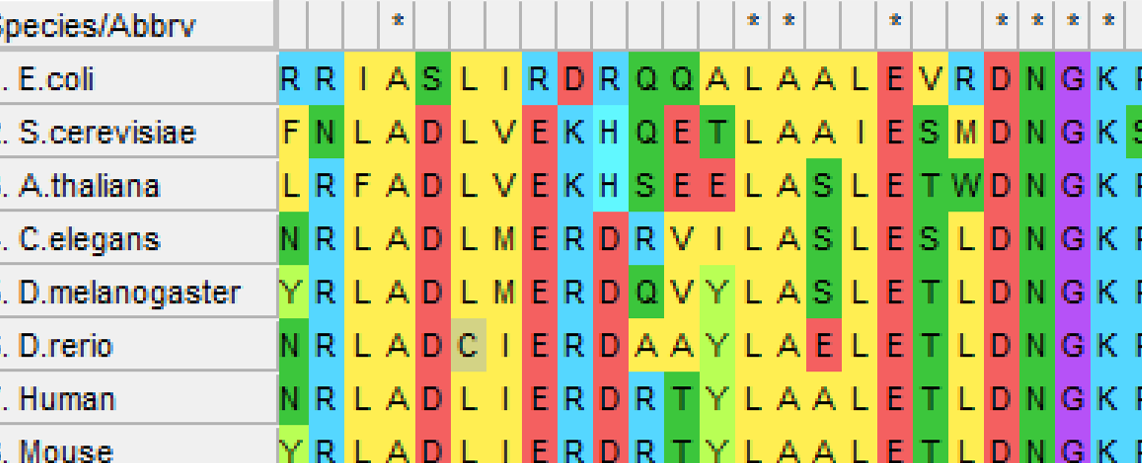
ALDH2 is a very highly conserved gene across multiple homologs. This protein has only one domain, aldedh, which is associated with the aldehyde dehydrogenase function. Mice have the closest protein to humans being 94.81% identical to humans.
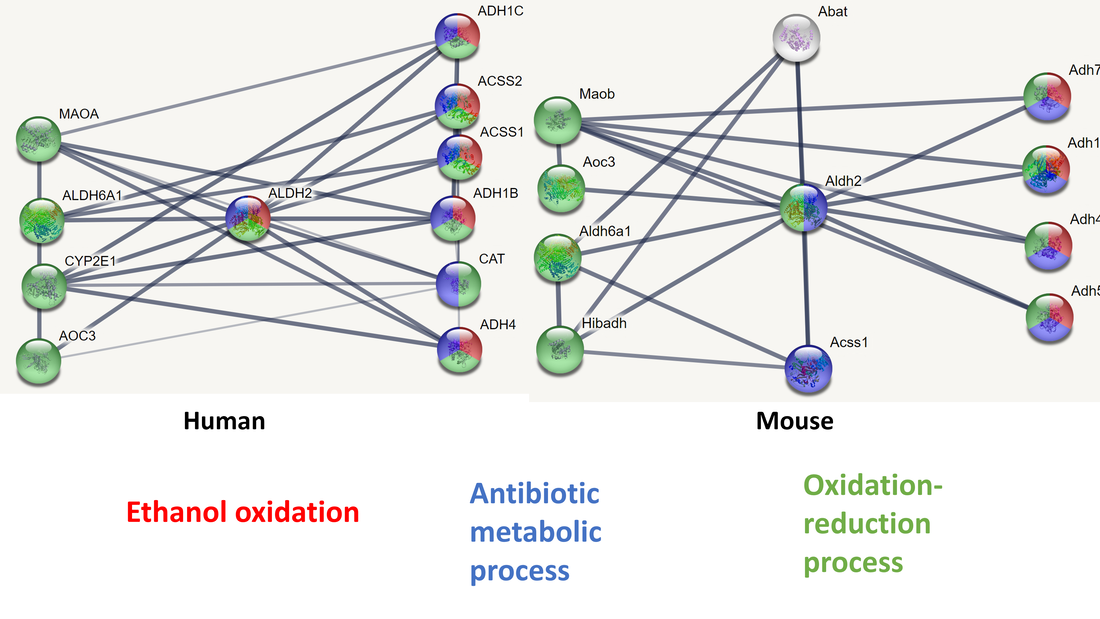
The protein interactions for ALDH2 show that most related proteins are involved in the oxidation-reduction process. In both mice and humans most interactive proteins appear to function somewhere in the metabolic process.
Gap in knowledge
It is still unknown how an ALDH2 deficiency contributes to a proliferative phenotype, but some attribute this to a increased risk of alcohol related cancer.
What model organism should I use?
I used mice as a model organism because mice have a very similar digestive systems, which means that alcohol will be processed in the same manner as humans. Also, multiple other studies have already created a mouse model with a knockout in this gene, so there are already methods to detect the functionality of this gene in a mouse model.
Aim #1:Determine conserved amino acids that are essential for ALDH2 function
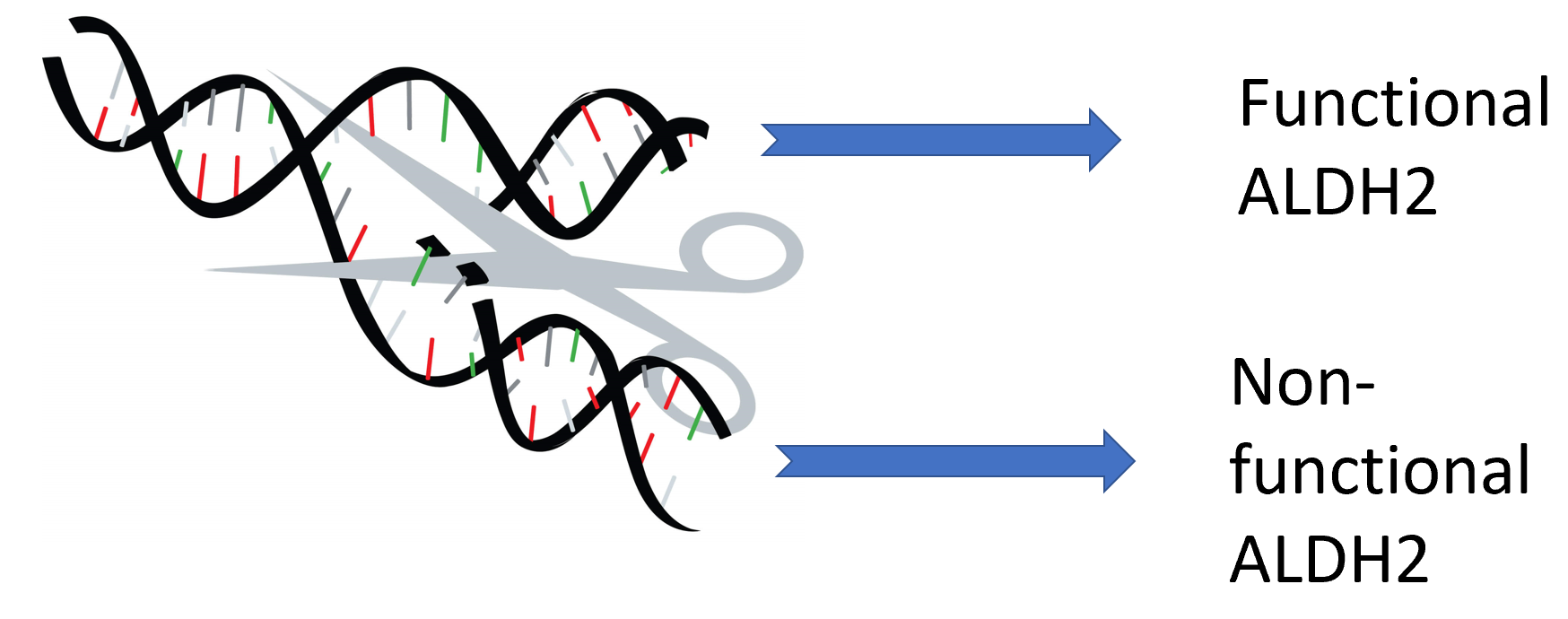
The goal of this aim is to identify conserved amino acids that are essential for normal ALDH2 activity. Using BLAST, I will identify homologs then align those sequences in ClustalOMEGA to determine highly conserved regions of the gene across species. Then, I would use CRISPR/Cas9 to create knockout mouse lines. Those mouse models would then be evaluated on their ability to process alcohol by testing the blood alcohol levels multiple times after alcohol consumption. I hypothesize that some knockout mutations will have a significantly metabolic rate.
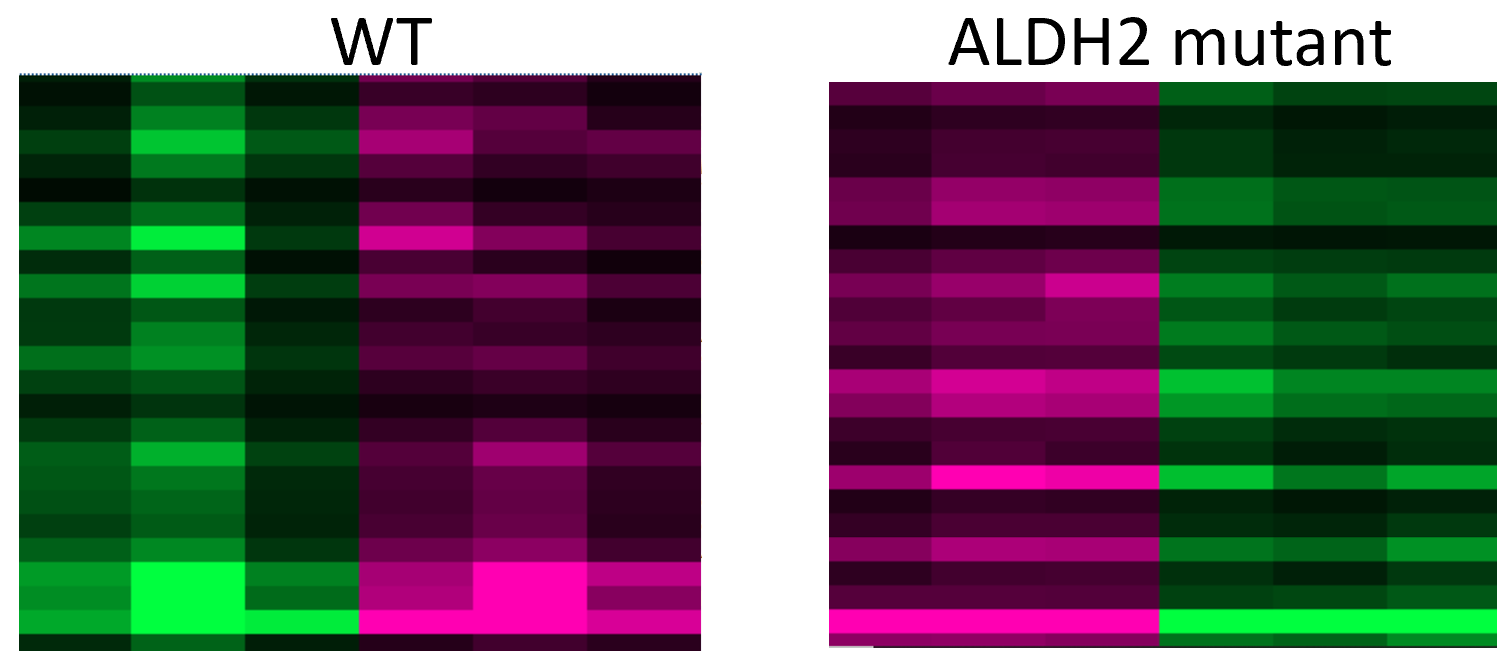
Aim #2: Identify differentially expressed genes in mutant ALDH2 mouse hepatocytes.
Using a mouse model with a E487K mutation the I will use RNA-seq to determine differentially expressed genes in mouse hepatocytes. I will extract liver cells from wild-type and mutant mice and will run RNA-seq on both types of cells. I expect to see some proliferative genes to be differentially expressed. Either oncogenes will be upregulated, or tumor suppressor genes will be down regulated. I expect that the mutant ALDH2 will cause the differential expression of these cells. Then, I will determine the proliferative pathway that causes the cancerous phenotype.
Aim #3:Characterize protein interactions that are important for normal liver development
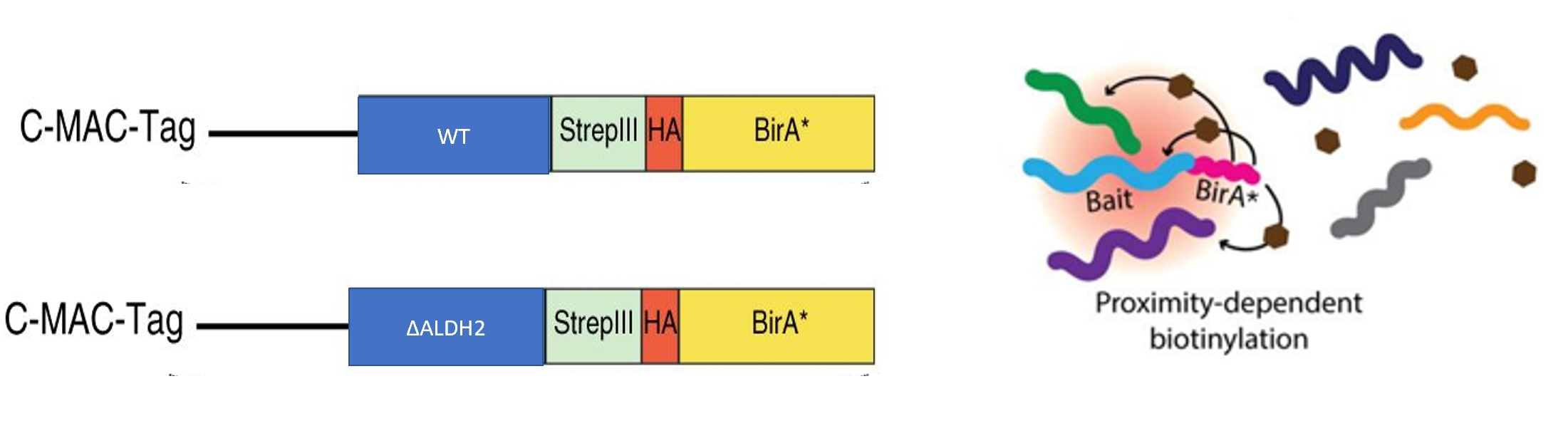
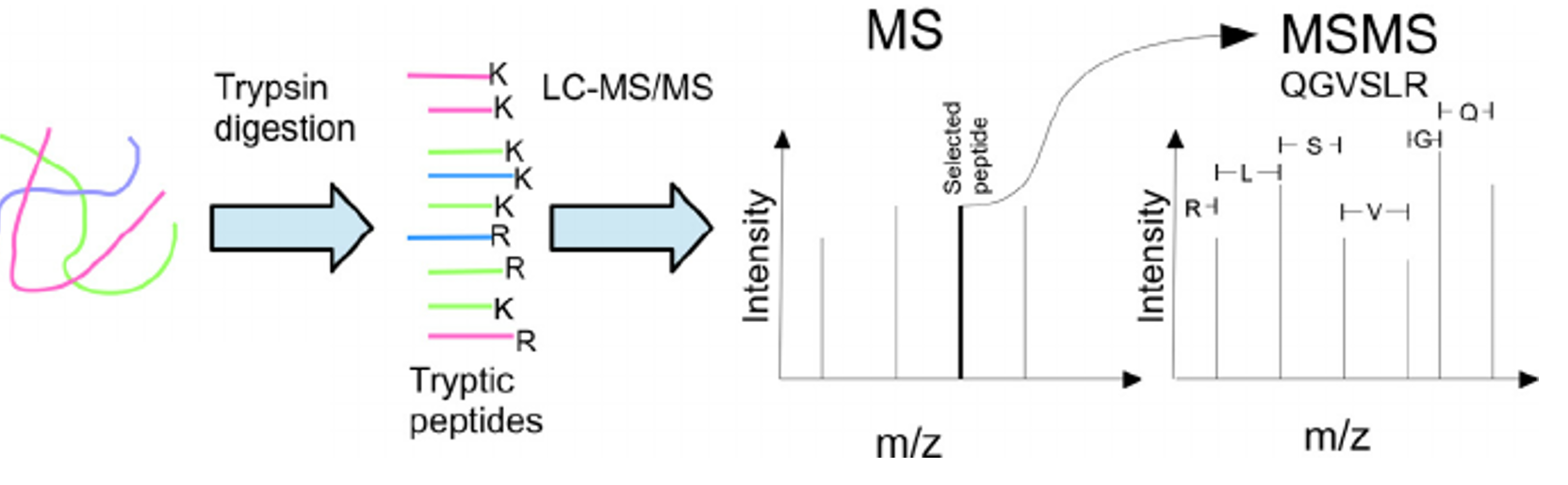
The mouse model from aim #2 will include a BirA gene attached to the mutant ALDH2 gene, so the BioID procedure can be performed. Then, mouse hepatocytes will be extracted from both groups of mice. These cells will then be exposed to an excess of biotin, so proteins nearby to ALDH2 will be biotinylated. The biotinylated proteins can be extracted using streptavin beads, and analyzed using trypsin digestion and tandem mass spectrometry. I can then determine the different interactions between wild-type and mutant ALDH2 proteins. I expect that mutant ALDH2 interacts with other proteins that are involved in controlling the cell cycle.
Future directions
After acquiring this data future experiments can be performed on other tissue types in the ALDH2 mutants. Head and neck tissues may be future focuses of how ALDH2 can cause cancer. These are two other tissue types in humans where ALDH2 human mutants commonly form cancerous cells.
Final presentation drafts
| Chris Stein aims draft 1.pptx | |
| File Size: | 5483 kb |
| File Type: | pptx |
| stein4-6.pptx | |
| File Size: | 7547 kb |
| File Type: | pptx |
| gen_564_final_presentation.pdf | |
| File Size: | 2931 kb |
| File Type: | |