What is transcriptonomics?
The transcriptome consists of the RNA molecules or transcripts within a cell (1). Studying the transcriptome provides insight into the product of transcription of genes. However, 95% of transcripts are never translated into proteins, which opens a new field of study into the activity of those RNA molecules. RNA molecules can function in other ways, like assisting in translation or interfering with other transcripts. The transcriptome is transcribed by genes in the genome, and is translated into proteins in the proteome. Transcriptome analysis provides insights into complex cellular processes that affect gene expression.
Methods to analyze the transcriptome
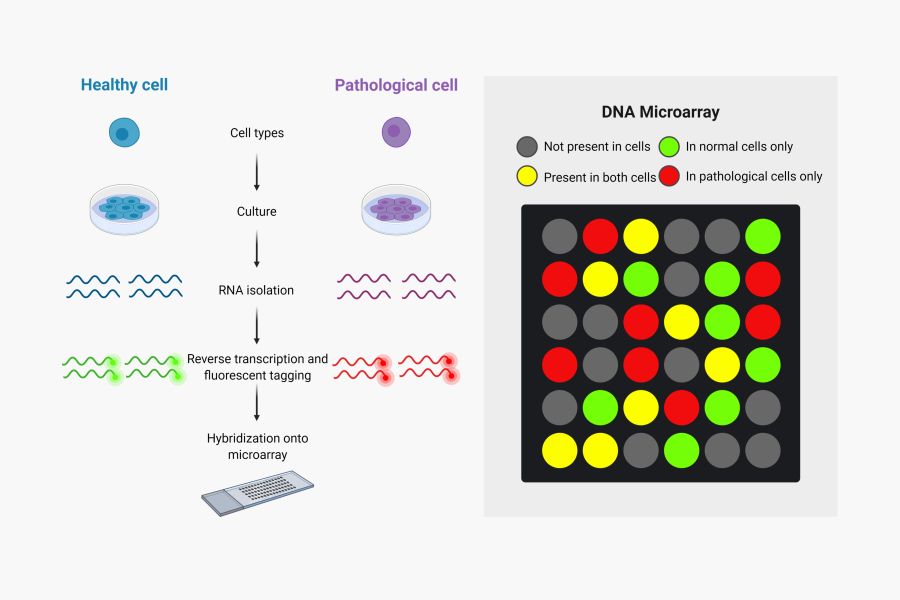
Microarray analysisMicroarrays are a transcriptomic technique that is uses to analyze a particular gene or set of genes (2). Microarrays use microscopic slides that have thousands of spots where tiny DNA molecules are placed (3). Many complementary DNA molecules (cDNA) are generated from an experimental and control lines (3). Then, the cDNAs are attached to fluorescent proteins. After being marked, the cDNA molecules are run across the microarray slide, which allows them to bind to their complementary strands (3).
|
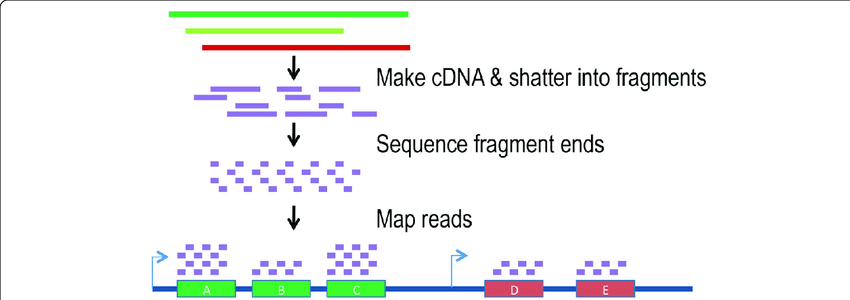
RNA sequencingRNA sequencing (RNA-seq) is a newer technology that takes advantage of next generation sequencing technologies (4). These procedures follow a common construct, which begins with RNA purification (5). After RNA purification, a DNA library needs to be formed, which may include reverse transcription. Then, sequencing can proceed, but in this case only fragments are sequenced and aligned against the genome (5). This gives an expression analysis of the majority of the genome.
|
Tissue expression of ALDH2
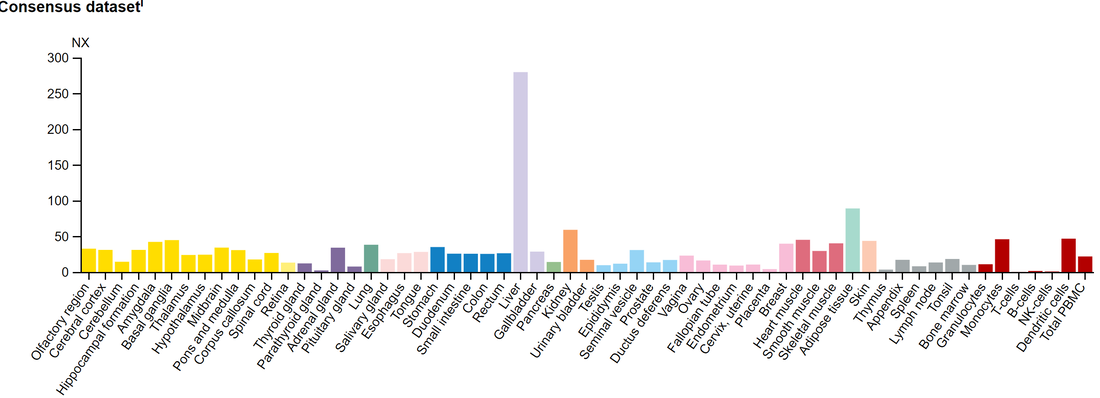
Figure 1: The tissues expression of ALDH2 provided by The Human Protein Atlas
Discussion
As expected, ALDH2 has high expression in the liver tissue. Most of the ethanol degradation pathway occurs in liver cells. There is also higher expression in adipose tissue, so this gene may be involved in oxidation of fatty acids.
References:
1)https://www.phgfoundation.org/blog/what-is-transcriptomics
2)https://www.nature.com/scitable/definition/microarray-202/
3)https://www.genome.gov/about-genomics/fact-sheets/DNA-Microarray-Technology
4)https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461
5)https://www.labome.com/method/RNA-seq.html
Pictures:
Header:https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3684953/
1)https://www.phgfoundation.org/blog/what-is-transcriptomics
2)https://www.nature.com/scitable/definition/microarray-202/
3)https://www.genome.gov/about-genomics/fact-sheets/DNA-Microarray-Technology
4)https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461
5)https://www.labome.com/method/RNA-seq.html
Pictures:
Header:https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3684953/